|
Thus, specific proteins lacking native tertiary structure, seem to be a universal precursor for GroEL-substrate binding. The most prominent feature associated with GroEL dependence is ( α/ β) 8 TIM (triosephosphate isomerase) barrel fold, albeit, it is also possessed by many cellular proteins which are GroEL independent folders. Although 250 cytosolic proteins interact with GroEL, ≈ 57 substrates have an obligate dependence on GroEL for folding. GroEL, an indispensable chaperone of Escherichia coli, selectively binds with unfolded proteins to assist in their productive folding. Several proteins fold spontaneously while others rely on molecular chaperones to reach their native states. 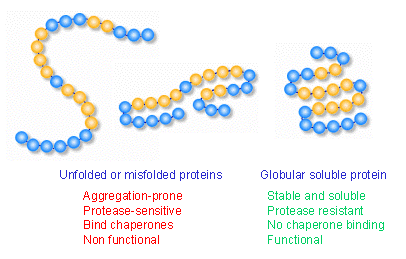
Properly folded proteins is a prerequisite in nearly every cellular process. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. SN and ST’s financial support is from CSIR network projects BSC0123 and BSC0124, respectively. KM is funded by IYBA fellowship from Department of Biotechnology. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are creditedĭata Availability: All relevant data are within the manuscript and associated Supporting Information files.įunding: LT is funded by INSPIRE Faculty Fellowship from Department of Science and Technology. Received: JanuAccepted: AugPublished: September 22, 2015Ĭopyright: © 2015 Nagpal et al. In summary, our work provides a molecular picture of an unfolded protein that is en route to chaperone binding, and these underlying structural properties might act as a molecular signal for their productive folding.Ĭitation: Nagpal S, Tiwari S, Mapa K, Thukral L (2015) Decoding Structural Properties of a Partially Unfolded Protein Substrate: En Route to Chaperone Binding. We also constructed network model of these intermediates that provides remarkable insights into stable hubs (or important residues) underlying diverse states of unfolded proteins. Significant increase in surface exposed hydrophobicity was observed in intermediates compared to native, which was further validated using ANS binding experiments. The most plausible intermediate was found to be in agreement with recently reported hydrogen-exchange experimental data. We use molecular dynamics simulations to reveal populated intermediate structures of DapA in atomic details. In this work, we study DapA protein, which exhibits obligate requirement on GroEL chaperonin machinery for its folding. However, structural information of non-folded proteins especially chaperone-dependent proteins is difficult to probe experimentally due to their inherent aggregation propensities. Many unfolded proteins if not folded accurately, become causal factors in various types of misfolding or aggregation induced diseases such as Alzheimer′s, Huntington′s and several other neurodegenerative disorders. 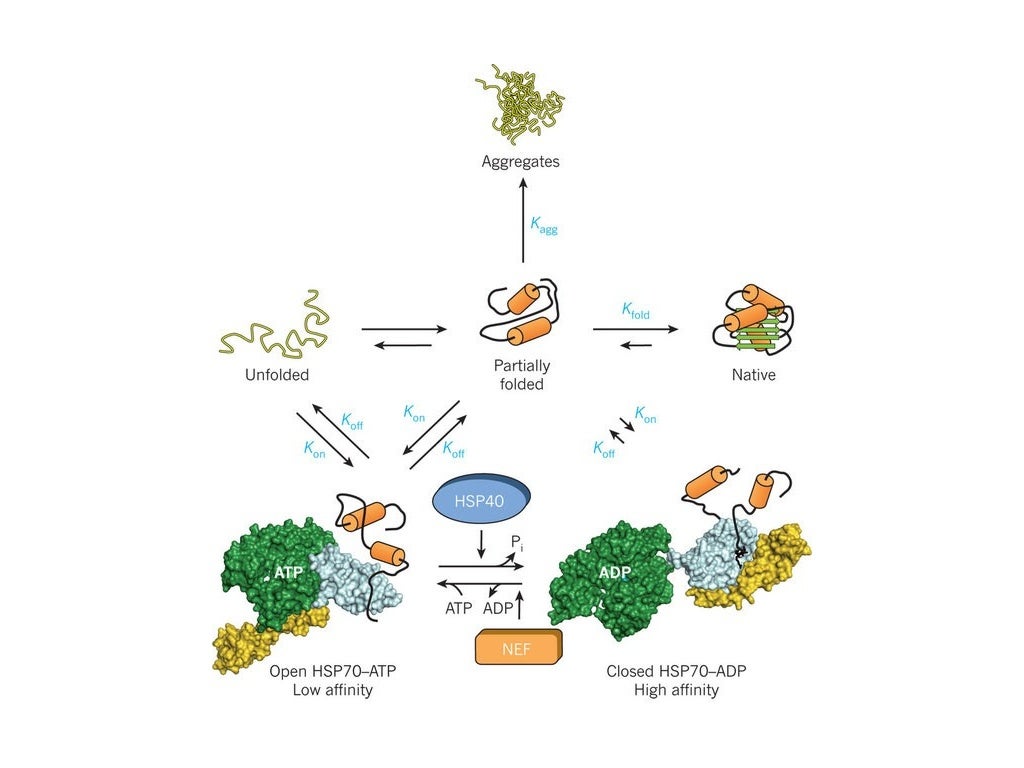
Several non-native proteins require molecular chaperones for proper folding. Overall, our results implicate that genomic variations (or mutations) in the distinct regions of protein structures might disrupt these topological signatures disabling chaperone-mediated folding, leading to formation of aggregates. Further, we constructed network graphs to elucidate long-range intra-protein connectivity of native and intermediate topologies, demonstrating regions that serve as central “hubs”. We validate the structural properties of these conformers using experimental data, including circular dichroism (CD), 1-anilinonaphthalene-8-sulfonic acid (ANS) binding measurements and previously reported hydrogen-deutrium exchange coupled to mass spectrometry (HDX-MS). We found that surface exposed hydrophobic patches are significantly increased, primarily contributed from native and non-native β-sheet elements. Thermodynamics analyses of unfolding simulations revealed populated intermediates with distinct structural characteristics. Here we performed all-atomistic and replica-exchange molecular dynamics simulations to dissect non-native ensemble of an obligate GroEL folder, DapA. Although the major structural components of GroEL are well defined, scaffolds of the non-native substrates that determine chaperone-mediated folding have been difficult to recognize. The E.coli chaperone, GroEL binds with a large number of unfolded and partially folded proteins, to facilitate proper folding and prevent misfolding and aggregation. Many proteins comprising of complex topologies require molecular chaperones to achieve their unique three-dimensional folded structure.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
